
| Name: PDZRN4 | Sequence: fasta or formatted (778aa) | NCBI GI: 142976783 | |
|
Description: PDZ domain containing RING finger 4
|
Referenced in: PDZ Domain
| ||
|
Composition:
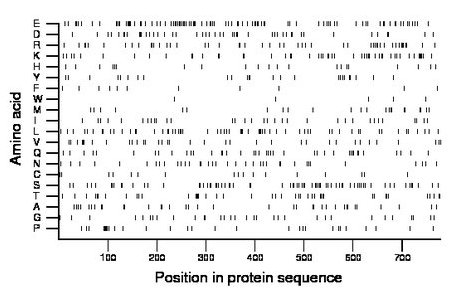
Amino acid Percentage Count Longest homopolymer A alanine 4.9 38 2 C cysteine 2.2 17 1 D aspartate 6.2 48 3 E glutamate 12.2 95 4 F phenylalanine 1.4 11 1 G glycine 4.2 33 2 H histidine 2.7 21 1 I isoleucine 4.4 34 2 K lysine 6.8 53 2 L leucine 8.2 64 2 M methionine 3.1 24 2 N asparagine 4.6 36 2 P proline 3.7 29 2 Q glutamine 5.1 40 2 R arginine 6.4 50 4 S serine 9.9 77 3 T threonine 5.5 43 2 V valine 4.5 35 2 W tryptophan 0.6 5 1 Y tyrosine 3.2 25 2 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 PDZ domain containing RING finger 4 PDZRN3 0.588 PDZ domain containing ring finger 3 PDZD4 0.426 PDZ domain containing 4 MPDZ 0.023 multiple PDZ domain protein INADL 0.023 InaD-like SCRIB 0.023 scribble isoform b SCRIB 0.023 scribble isoform a SYNJ2BP 0.022 synaptojanin 2 binding protein PARD3B 0.020 par-3 partitioning defective 3 homolog B isoform b ... LNX2 0.020 ligand of numb-protein X 2 IL16 0.020 interleukin 16 isoform 2 PARD3B 0.020 par-3 partitioning defective 3 homolog B isoform a ... PARD3B 0.020 par-3 partitioning defective 3 homolog B isoform c ... DLG2 0.018 chapsyn-110 isoform 3 DLG2 0.018 chapsyn-110 isoform 1 DLG2 0.018 chapsyn-110 isoform 2 DLG1 0.017 discs, large homolog 1 isoform 1 DLG1 0.017 discs, large homolog 1 isoform 2 DFNB31 0.016 CASK-interacting protein CIP98 isoform 1 PDZD2 0.016 PDZ domain containing 2 LNX1 0.016 ligand of numb-protein X 1 isoform a LNX1 0.016 ligand of numb-protein X 1 isoform b GCC2 0.016 GRIP and coiled-coil domain-containing 2 DLG4 0.016 post-synaptic density protein 95 isoform 2 DLG4 0.016 post-synaptic density protein 95 isoform 1 MYST3 0.015 MYST histone acetyltransferase (monocytic leukemia)... MYST3 0.015 MYST histone acetyltransferase (monocytic leukemia)... MYST3 0.015 MYST histone acetyltransferase (monocytic leukemia)... SLK 0.014 serine/threonine kinase 2 DVL1 0.014 dishevelled 1Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
