
| Name: LOC100128256 | Sequence: fasta or formatted (240aa) | NCBI GI: 169212746 | |
|
Description: PREDICTED: hypothetical protein
| Not currently referenced in the text | ||
Other entries for this name:
alt mRNA {240aa} PREDICTED: hypothetical protein | |||
|
Composition:
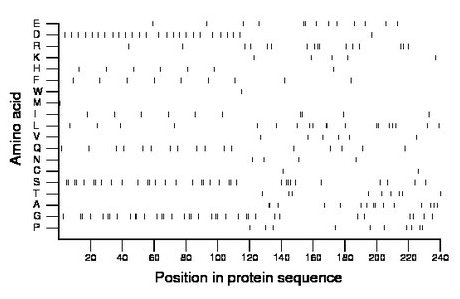
Amino acid Percentage Count Longest homopolymer A alanine 6.2 15 2 C cysteine 0.8 2 1 D aspartate 10.8 26 1 E glutamate 5.4 13 2 F phenylalanine 3.8 9 1 G glycine 15.0 36 2 H histidine 2.9 7 1 I isoleucine 4.2 10 2 K lysine 2.1 5 1 L leucine 7.9 19 2 M methionine 0.4 1 1 N asparagine 1.7 4 1 P proline 4.2 10 1 Q glutamine 7.1 17 1 R arginine 6.7 16 2 S serine 14.2 34 2 T threonine 3.8 9 1 V valine 2.5 6 1 W tryptophan 0.4 1 1 Y tyrosine 0.0 0 0 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 PREDICTED: hypothetical protein LOC100128256 1.000 PREDICTED: hypothetical protein TAF15 0.054 TBP-associated factor 15 isoform 2 TAF15 0.054 TBP-associated factor 15 isoform 1 DMKN 0.037 dermokine isoform 4 precursor DMKN 0.037 dermokine isoform 5 precursor DMKN 0.037 dermokine isoform 3 precursor DMKN 0.037 dermokine isoform 2 precursor LOC100286952 0.033 PREDICTED: hypothetical protein XP_002344252 DSPP 0.031 dentin sialophosphoprotein preproprotein LEO1 0.031 Leo1, Paf1/RNA polymerase II complex component, homo... KRT10 0.031 keratin 10 KRT24 0.028 keratin 24 RBMXL3 0.026 RNA binding motif protein, X-linked-like 3 ACRC 0.024 ACRC protein HCLS1 0.022 hematopoietic cell-specific Lyn substrate 1 FUS 0.022 fusion (involved in t(12;16) in malignant liposarcoma... TNRC18 0.022 trinucleotide repeat containing 18 IRS4 0.020 insulin receptor substrate 4 KRT9 0.020 keratin 9 FAM98A 0.020 hypothetical protein LOC25940 FLG 0.017 filaggrin PMS2 0.017 PMS2 postmeiotic segregation increased 2 isoform a [H... KRT2 0.017 keratin 2 LOR 0.017 loricrin LOC100128084 0.015 PREDICTED: hypothetical protein FSCB 0.015 fibrous sheath CABYR binding protein FMN1 0.013 formin 1 HNRNPA1 0.013 heterogeneous nuclear ribonucleoprotein A1 isoform b... HNRNPA3 0.013 heterogeneous nuclear ribonucleoprotein A3Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
