
| Name: TGM4 | Sequence: fasta or formatted (684aa) | NCBI GI: 156627577 | |
|
Description: transglutaminase 4 (prostate)
|
Referenced in:
| ||
|
Composition:
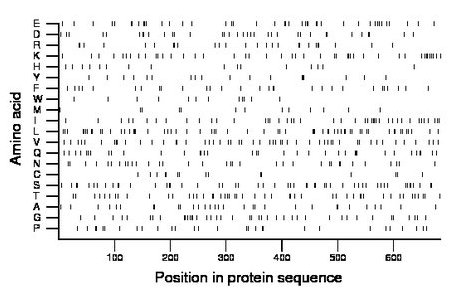
Amino acid Percentage Count Longest homopolymer A alanine 4.5 31 2 C cysteine 2.2 15 2 D aspartate 5.1 35 2 E glutamate 6.1 42 2 F phenylalanine 4.2 29 1 G glycine 6.0 41 2 H histidine 2.5 17 2 I isoleucine 6.3 43 2 K lysine 6.4 44 2 L leucine 9.1 62 3 M methionine 2.2 15 2 N asparagine 5.0 34 1 P proline 4.2 29 1 Q glutamine 5.3 36 2 R arginine 3.9 27 3 S serine 7.6 52 2 T threonine 7.2 49 2 V valine 7.6 52 2 W tryptophan 1.8 12 1 Y tyrosine 2.8 19 2 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary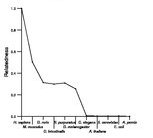 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 transglutaminase 4 (prostate) TGM1 0.330 transglutaminase 1 TGM7 0.289 transglutaminase 7 TGM6 0.289 transglutaminase 6 F13A1 0.273 coagulation factor XIII A1 subunit precursor TGM3 0.267 transglutaminase 3 precursor TGM5 0.264 transglutaminase 5 isoform 1 TGM2 0.245 transglutaminase 2 isoform a TGM2 0.233 transglutaminase 2 isoform b TGM5 0.226 transglutaminase 5 isoform 2 EPB42 0.177 erythrocyte membrane protein band 4.2 isoform 1 [Ho... EPB42 0.176 erythrocyte membrane protein band 4.2 isoform 2 [Ho... PCDHA5 0.004 protocadherin alpha 5 isoform 1 precursor PCDHA5 0.004 protocadherin alpha 5 isoform 2 precursor ATP2A3 0.004 ATPase, Ca++ transporting, ubiquitous isoform a [Hom... ATP2A3 0.004 ATPase, Ca++ transporting, ubiquitous isoform c [Hom... ATP2A3 0.004 ATPase, Ca++ transporting, ubiquitous isoform f [Hom... ATP2A3 0.004 ATPase, Ca++ transporting, ubiquitous isoform c [Hom... ATP2A3 0.004 ATPase, Ca++ transporting, ubiquitous isoform b [Hom... ATP2A3 0.004 ATPase, Ca++ transporting, ubiquitous isoform d [Hom... ATP2A3 0.004 ATPase, Ca++ transporting, ubiquitous isoform e [Hom... MAPK15 0.004 mitogen-activated protein kinase 15 XCR1 0.004 XC chemokine receptor 1 XCR1 0.004 XC chemokine receptor 1 EPHA1 0.004 ephrin receptor EphA1 C1orf226 0.004 hypothetical protein LOC400793 isoform 2 C1orf226 0.004 hypothetical protein LOC400793 isoform 1 BYSL 0.004 bystin CYTL1 0.004 cytokine-like 1 PKD1L1 0.004 polycystin-1L1Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
