
| Name: POU3F3 | Sequence: fasta or formatted (500aa) | NCBI GI: 5453936 | |
|
Description: POU class 3 homeobox 3
|
Referenced in:
| ||
|
Composition:
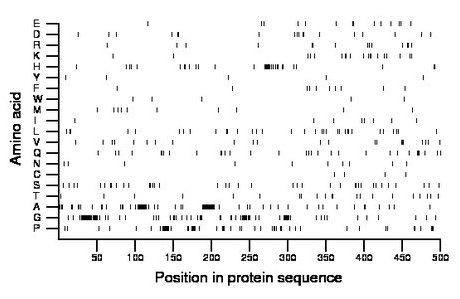
Amino acid Percentage Count Longest homopolymer A alanine 14.0 70 16 C cysteine 0.8 4 1 D aspartate 3.8 19 2 E glutamate 2.8 14 2 F phenylalanine 2.0 10 1 G glycine 15.8 79 14 H histidine 6.6 33 8 I isoleucine 1.6 8 1 K lysine 3.8 19 2 L leucine 6.6 33 2 M methionine 2.2 11 1 N asparagine 2.0 10 1 P proline 10.8 54 6 Q glutamine 6.2 31 3 R arginine 2.8 14 2 S serine 7.6 38 2 T threonine 4.2 21 2 V valine 4.4 22 2 W tryptophan 1.0 5 1 Y tyrosine 1.0 5 1 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary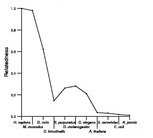 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 POU class 3 homeobox 3 POU3F2 0.416 POU domain, class 3, transcription factor 2 POU3F4 0.344 POU domain, class 3, transcription factor 4 POU3F1 0.337 POU domain, class 3, transcription factor 1 POU2F1 0.177 POU class 2 homeobox 1 POU2F3 0.173 POU transcription factor POU2F2 0.165 POU domain, class 2, transcription factor 2 POU5F1 0.161 POU domain, class 5, transcription factor 1 isoform ... POU5F1 0.160 POU domain, class 5, transcription factor 1 isoform... POU5F1B 0.154 POU class 5 homeobox 1B POU1F1 0.152 pituitary specific transcription factor 1 isoform b... POU1F1 0.152 pituitary specific transcription factor 1 isoform alp... POU4F1 0.151 POU domain, class 4, transcription factor 1 POU4F2 0.150 Brn3b POU domain transcription factor POU4F3 0.126 POU class 4 homeobox 3 POU5F2 0.114 POU domain class 5, transcription factor 2 POU6F1 0.108 POU class 6 homeobox 1 POU6F2 0.078 POU domain, class 6, transcription factor 2 ONECUT3 0.061 one cut homeobox 3 ARID1B 0.056 AT rich interactive domain 1B (SWI1-like) isoform 2 ... ARID1B 0.056 AT rich interactive domain 1B (SWI1-like) isoform 1 ... ARID1B 0.056 AT rich interactive domain 1B (SWI1-like) isoform 3 ... PRR12 0.055 proline rich 12 GATA6 0.054 GATA binding protein 6 FUSSEL18 0.054 PREDICTED: functional smad suppressing element 18 [... PPP1R10 0.053 protein phosphatase 1, regulatory subunit 10 FUSSEL18 0.053 PREDICTED: functional smad suppressing element 18 [... FUSSEL18 0.053 PREDICTED: functional smad suppressing element 18 [... SFPQ 0.044 splicing factor proline/glutamine rich (polypyrimidin... LBXCOR1 0.040 LBXCOR1 homologHuman BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
