
| Name: MRPL49 | Sequence: fasta or formatted (166aa) | NCBI GI: 4826649 | |
|
Description: mitochondrial ribosomal protein L49
|
Referenced in:
| ||
|
Composition:
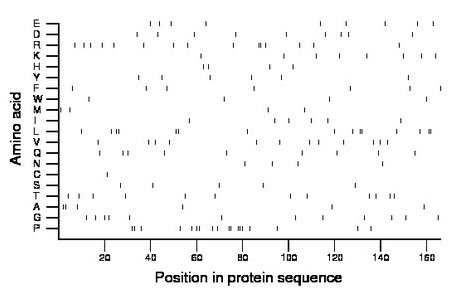
Amino acid Percentage Count Longest homopolymer A alanine 3.6 6 2 C cysteine 0.6 1 1 D aspartate 5.4 9 1 E glutamate 5.4 9 1 F phenylalanine 4.2 7 1 G glycine 7.2 12 1 H histidine 2.4 4 1 I isoleucine 3.6 6 1 K lysine 4.8 8 1 L leucine 9.0 15 2 M methionine 2.4 4 1 N asparagine 2.4 4 1 P proline 10.8 18 3 Q glutamine 5.4 9 1 R arginine 9.0 15 2 S serine 3.0 5 1 T threonine 7.2 12 1 V valine 7.2 12 1 W tryptophan 2.4 4 1 Y tyrosine 3.6 6 1 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 mitochondrial ribosomal protein L49 LILRB5 0.024 leukocyte immunoglobulin-like receptor, subfamily B... KDM6B 0.024 lysine (K)-specific demethylase 6B HLF 0.021 hepatic leukemia factor SFPQ 0.021 splicing factor proline/glutamine rich (polypyrimidin... C1orf88 0.018 hypothetical protein LOC128344 MLXIPL 0.018 Williams Beuren syndrome chromosome region 14 isofor... MLXIPL 0.018 Williams Beuren syndrome chromosome region 14 isofor... MLXIPL 0.018 Williams Beuren syndrome chromosome region 14 isofor... MLXIPL 0.018 Williams Beuren syndrome chromosome region 14 isofor... EFS 0.018 embryonal Fyn-associated substrate isoform 2 EFS 0.018 embryonal Fyn-associated substrate isoform 1 LMOD2 0.018 leiomodin 2 (cardiac) RUSC1 0.015 RUN and SH3 domain containing 1 isoform b RUSC1 0.015 RUN and SH3 domain containing 1 isoform a LOC100128370 0.015 PREDICTED: hypothetical protein LOC100128370 0.015 PREDICTED: hypothetical protein MYO3B 0.015 myosin IIIB isoform 1 MYO3B 0.015 myosin IIIB isoform 2 GCC2 0.015 GRIP and coiled-coil domain-containing 2 PSORS1C2 0.015 SPR1 protein WWP1 0.012 WW domain containing E3 ubiquitin protein ligase 1 [... POU3F1 0.012 POU domain, class 3, transcription factor 1 TTN 0.012 titin isoform N2-A DIAPH1 0.012 diaphanous 1 isoform 2 DIAPH1 0.012 diaphanous 1 isoform 1 WASF2 0.012 WAS protein family, member 2 PRB3 0.012 proline-rich protein BstNI subfamily 3 precursor [H... WBP11 0.012 WW domain binding protein 11 SRRM1 0.012 serine/arginine repetitive matrix 1Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
