
| Name: EVX1 | Sequence: fasta or formatted (407aa) | NCBI GI: 4503615 | |
|
Description: even-skipped homeobox 1
|
Referenced in:
| ||
|
Composition:
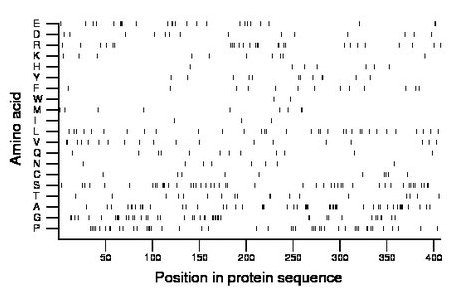
Amino acid Percentage Count Longest homopolymer A alanine 12.3 50 4 C cysteine 2.5 10 1 D aspartate 3.4 14 1 E glutamate 6.1 25 3 F phenylalanine 3.2 13 1 G glycine 10.8 44 4 H histidine 1.7 7 1 I isoleucine 0.7 3 1 K lysine 2.5 10 1 L leucine 8.1 33 2 M methionine 2.2 9 2 N asparagine 2.0 8 1 P proline 11.1 45 2 Q glutamine 2.9 12 1 R arginine 6.6 27 2 S serine 11.3 46 3 T threonine 4.4 18 2 V valine 4.7 19 2 W tryptophan 0.5 2 1 Y tyrosine 2.9 12 2 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 even-skipped homeobox 1 EVX2 0.390 even-skipped homeobox 2 HOXB3 0.095 homeobox B3 NKX1-1 0.077 PREDICTED: HPX-153 homeobox NKX1-1 0.076 PREDICTED: NK1 homeobox 1 ARX 0.076 aristaless related homeobox GBX1 0.076 gastrulation brain homeo box 1 HOXB2 0.074 homeobox B2 VAX2 0.072 ventral anterior homeobox 2 HOXA3 0.071 homeobox A3 isoform a HOXA3 0.071 homeobox A3 isoform a HOXD3 0.071 homeobox D3 HMX1 0.067 homeo box (H6 family) 1 NKX3-2 0.067 NK3 homeobox 2 HOXA2 0.066 homeobox A2 HOXA3 0.066 homeobox A3 isoform b PDX1 0.064 pancreatic and duodenal homeobox 1 MEOX2 0.064 mesenchyme homeobox 2 NKX1-1 0.064 PREDICTED: NK1 homeobox 1 GBX2 0.064 gastrulation brain homeo box 2 BARHL2 0.063 BarH-like homeobox 2 HMX3 0.063 H6 family homeobox 3 BARHL1 0.063 BarH-like homeobox 1 VAX1 0.063 ventral anterior homeobox 1 isoform a HOXB1 0.062 homeobox B1 CDX1 0.062 caudal type homeobox 1 NKX2-4 0.061 NK2 homeobox 4 HOXD4 0.059 homeobox D4 MNX1 0.059 homeo box HB9 HOXD11 0.059 homeobox D11Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
