
| Name: NRXN2 | Sequence: fasta or formatted (666aa) | NCBI GI: 21166382 | |
|
Description: neurexin 2 isoform beta precursor
|
Referenced in:
| ||
Other entries for this name:
alt prot [1712aa] neurexin 2 isoform alpha-1 precursor alt prot [1642aa] neurexin 2 isoform alpha-2 precursor | |||
|
Composition:
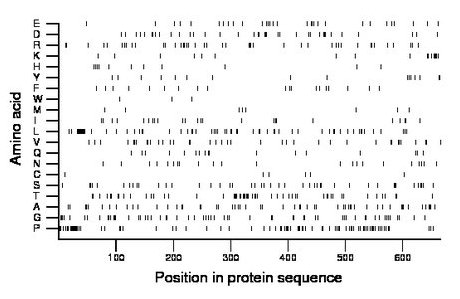
Amino acid Percentage Count Longest homopolymer A alanine 8.6 57 4 C cysteine 0.9 6 1 D aspartate 5.7 38 2 E glutamate 4.7 31 2 F phenylalanine 2.9 19 1 G glycine 9.2 61 2 H histidine 2.0 13 2 I isoleucine 3.6 24 2 K lysine 2.6 17 2 L leucine 9.5 63 7 M methionine 1.4 9 1 N asparagine 3.3 22 1 P proline 12.6 84 10 Q glutamine 2.6 17 1 R arginine 6.6 44 2 S serine 6.8 45 3 T threonine 8.6 57 4 V valine 5.9 39 2 W tryptophan 0.5 3 1 Y tyrosine 2.6 17 2 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 neurexin 2 isoform beta precursor NRXN2 0.858 neurexin 2 isoform alpha-1 precursor NRXN2 0.800 neurexin 2 isoform alpha-2 precursor NRXN1 0.310 neurexin 1 isoform alpha2 precursor NRXN1 0.295 neurexin 1 isoform beta precursor NRXN3 0.295 neurexin 3 isoform 3 precursor NRXN3 0.258 neurexin 3 isoform 2 precursor NRXN1 0.255 neurexin 1 isoform alpha1 precursor NRXN3 0.227 neurexin 3 isoform 1 precursor PRR12 0.024 proline rich 12 FLJ22184 0.022 PREDICTED: hypothetical protein FLJ22184 FLJ22184 0.022 PREDICTED: hypothetical protein FLJ22184 FLJ22184 0.022 PREDICTED: hypothetical protein LOC80164 TENC1 0.021 tensin like C1 domain containing phosphatase isoform... TENC1 0.021 tensin like C1 domain containing phosphatase isoform... TENC1 0.021 tensin like C1 domain containing phosphatase isoform... SYN1 0.020 synapsin I isoform Ia SYN1 0.020 synapsin I isoform Ib CEL 0.019 carboxyl ester lipase precursor ZC3H4 0.019 zinc finger CCCH-type containing 4 MUC2 0.019 mucin 2 precursor HCN4 0.017 hyperpolarization activated cyclic nucleotide-gated p... PRG4 0.017 proteoglycan 4 isoform D PRG4 0.017 proteoglycan 4 isoform C PRG4 0.017 proteoglycan 4 isoform B PRG4 0.017 proteoglycan 4 isoform A C2orf55 0.017 hypothetical protein LOC343990 RNF215 0.017 ring finger protein 215 COL1A1 0.016 alpha 1 type I collagen preproprotein COL5A1 0.016 alpha 1 type V collagen preproproteinHuman BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
