
| Name: FERMT2 | Sequence: fasta or formatted (680aa) | NCBI GI: 29789006 | |
|
Description: fermitin family homolog 2 isoform 1
|
Referenced in: Cytoskeleton
| ||
Other entries for this name:
alt prot [687aa] fermitin family homolog 2 isoform 2 alt prot [633aa] fermitin family homolog 2 isoform 3 | |||
|
Composition:
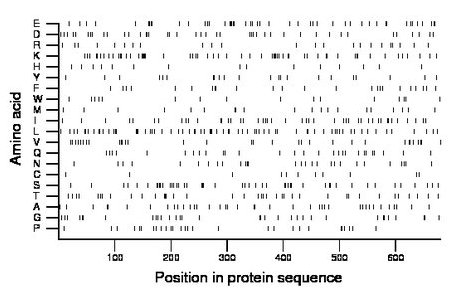
Amino acid Percentage Count Longest homopolymer A alanine 6.0 41 2 C cysteine 1.6 11 1 D aspartate 6.2 42 2 E glutamate 7.4 50 3 F phenylalanine 3.5 24 2 G glycine 4.9 33 2 H histidine 2.2 15 1 I isoleucine 6.6 45 1 K lysine 9.3 63 6 L leucine 10.3 70 3 M methionine 3.2 22 3 N asparagine 4.1 28 2 P proline 4.1 28 1 Q glutamine 4.1 28 1 R arginine 3.4 23 1 S serine 6.9 47 2 T threonine 5.7 39 2 V valine 4.7 32 2 W tryptophan 2.5 17 2 Y tyrosine 3.2 22 2 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 fermitin family homolog 2 isoform 1 FERMT2 0.995 fermitin family homolog 2 isoform 2 FERMT2 0.912 fermitin family homolog 2 isoform 3 FERMT1 0.631 kindlin-1 FERMT3 0.525 fermitin family homolog 3 short form FERMT3 0.521 fermitin family homolog 3 long form TLN1 0.033 talin 1 TLN2 0.028 talin 2 AFAP1L2 0.011 KIAA1914 protein isoform 1 AFAP1L2 0.011 KIAA1914 protein isoform 2 AFAP1 0.009 actin filament associated protein 1 isoform a [Homo... AFAP1 0.009 actin filament associated protein 1 OPA1 0.008 optic atrophy 1 isoform 8 OPA1 0.008 optic atrophy 1 isoform 7 OPA1 0.008 optic atrophy 1 isoform 6 OPA1 0.008 optic atrophy 1 isoform 5 OPA1 0.008 optic atrophy 1 isoform 4 OPA1 0.008 optic atrophy 1 isoform 3 OPA1 0.008 optic atrophy 1 isoform 2 OPA1 0.008 optic atrophy 1 isoform 1 TOPORS 0.007 topoisomerase I binding, arginine/serine-rich RDX 0.006 radixin POLN 0.005 DNA-directed DNA polymerase nu OSBP 0.005 oxysterol binding protein EPB41L3 0.004 erythrocyte membrane protein band 4.1-like 3 RAD18 0.004 postreplication repair protein hRAD18p MYH10 0.004 myosin, heavy polypeptide 10, non-muscle OSBPL8 0.004 oxysterol-binding protein-like protein 8 isoform b [... OSBPL8 0.004 oxysterol-binding protein-like protein 8 isoform a [... HOMER1 0.004 homer 1Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
