
| Name: MCFD2 | Sequence: fasta or formatted (146aa) | NCBI GI: 21281683 | |
|
Description: multiple coagulation factor deficiency 2
|
Referenced in: Calmodulin and Calcium
| ||
|
Composition:
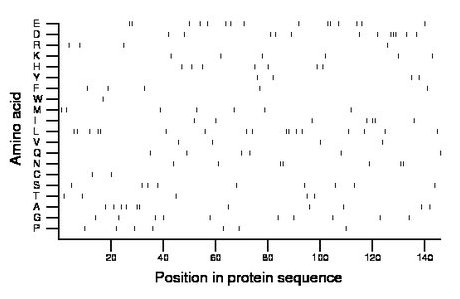
Amino acid Percentage Count Longest homopolymer A alanine 7.5 11 2 C cysteine 1.4 2 1 D aspartate 8.2 12 3 E glutamate 10.3 15 2 F phenylalanine 3.4 5 1 G glycine 6.8 10 1 H histidine 4.8 7 1 I isoleucine 4.8 7 2 K lysine 4.1 6 1 L leucine 11.6 17 2 M methionine 4.8 7 1 N asparagine 4.8 7 2 P proline 4.8 7 1 Q glutamine 4.1 6 1 R arginine 2.7 4 1 S serine 6.2 9 1 T threonine 3.4 5 1 V valine 2.7 4 1 W tryptophan 0.7 1 1 Y tyrosine 2.7 4 1 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 multiple coagulation factor deficiency 2 CGREF1 0.076 cell growth regulator with EF-hand domain 1 CIB2 0.072 DNA-dependent protein kinase catalytic subunit-intera... CIB4 0.058 calcium and integrin binding family member 4 GUCA1B 0.051 guanylate cyclase activator 1B (retina) GUCA1A 0.047 guanylate cyclase activator 1A (retina) GUCA1C 0.047 guanylate cyclase activator 1C TNNC2 0.043 fast skeletal muscle troponin C CHP2 0.043 hepatocellular carcinoma antigen gene 520 NUCB2 0.043 nucleobindin 2 CIB1 0.043 calcium and integrin binding 1 TNNC1 0.036 troponin C, slow EFHB 0.036 EF hand domain family, member B HPCAL1 0.036 hippocalcin-like 1 HPCAL1 0.036 hippocalcin-like 1 CALB2 0.032 calbindin 2 isoform 1 CALB2 0.032 calbindin 2 isoform 22k CALM3 0.032 calmodulin 3 CALML3 0.032 calmodulin-like 3 CALM2 0.032 calmodulin 2 CALM1 0.032 calmodulin 1 VSNL1 0.032 visinin-like 1 KCNIP1 0.032 Kv channel interacting protein 1 isoform 3 KCNIP1 0.032 Kv channel interacting protein 1 isoform 1 KCNIP1 0.032 Kv channel interacting protein 1 isoform 2 NUCB1 0.029 nucleobindin 1 HPCA 0.029 hippocalcin HPCAL4 0.025 hippocalcin-like protein 4 CIT 0.025 citron CALML5 0.022 calmodulin-like 5Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
