
| Name: NEO1 | Sequence: fasta or formatted (1461aa) | NCBI GI: 157311649 | |
|
Description: neogenin homolog 1
|
Referenced in: Fibronectin Family
| ||
|
Composition:
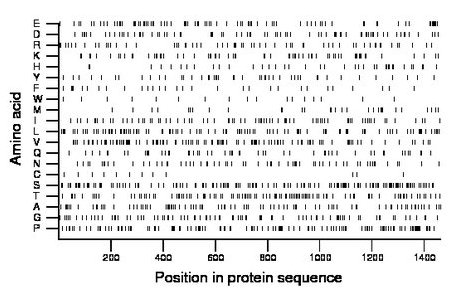
Amino acid Percentage Count Longest homopolymer A alanine 6.4 93 4 C cysteine 0.9 13 2 D aspartate 5.1 75 2 E glutamate 5.5 80 2 F phenylalanine 2.1 30 1 G glycine 6.0 87 2 H histidine 2.9 42 3 I isoleucine 5.3 78 2 K lysine 4.7 68 3 L leucine 7.3 107 4 M methionine 2.2 32 3 N asparagine 4.4 64 2 P proline 8.9 130 3 Q glutamine 3.6 52 2 R arginine 4.5 66 2 S serine 10.1 148 3 T threonine 7.7 113 2 V valine 7.9 115 4 W tryptophan 1.2 17 1 Y tyrosine 3.5 51 2 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 neogenin homolog 1 DCC 0.477 deleted in colorectal carcinoma PRTG 0.089 protogenin PTPRD 0.084 protein tyrosine phosphatase, receptor type, D isofo... IGDCC4 0.084 immunoglobulin superfamily, DCC subclass, member 4 [... PTPRS 0.083 protein tyrosine phosphatase, receptor type, sigma ... PTPRD 0.082 protein tyrosine phosphatase, receptor type, D isofor... PTPRS 0.080 protein tyrosine phosphatase, receptor type, sigma ... PTPRD 0.078 protein tyrosine phosphatase, receptor type, D isofo... PTPRF 0.076 protein tyrosine phosphatase, receptor type, F isof... DSCAM 0.076 Down syndrome cell adhesion molecule isoform CHD2-42... PTPRF 0.075 protein tyrosine phosphatase, receptor type, F isof... DSCAML1 0.075 Down syndrome cell adhesion molecule like 1 SDK2 0.074 sidekick 2 IGDCC3 0.074 putative neuronal cell adhesion molecule ROBO2 0.072 roundabout, axon guidance receptor, homolog 2 isofo... ROBO2 0.072 roundabout, axon guidance receptor, homolog 2 isofor... SDK1 0.070 sidekick 1 L1CAM 0.070 L1 cell adhesion molecule isoform 3 precursor [Homo... L1CAM 0.070 L1 cell adhesion molecule isoform 2 precursor L1CAM 0.070 L1 cell adhesion molecule isoform 1 precursor NRCAM 0.070 neuronal cell adhesion molecule isoform A precursor ... CNTN4 0.069 contactin 4 isoform a precursor NRCAM 0.069 neuronal cell adhesion molecule isoform B precursor ... ROBO1 0.068 roundabout 1 isoform b ROBO1 0.068 roundabout 1 isoform a CNTN3 0.067 contactin 3 ROBO1 0.066 roundabout 1 isoform d ROBO1 0.066 roundabout 1 isoform c LOC642132 0.066 PREDICTED: similar to roundabout 1 isoform 1Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
