
| Name: VPS54 | Sequence: fasta or formatted (977aa) | NCBI GI: 54234034 | |
|
Description: vacuolar protein sorting 54 isoform 1
|
Referenced in: ER, Golgi, and the Secretory Pathway
| ||
Other entries for this name:
alt prot [965aa] vacuolar protein sorting 54 isoform 2 | |||
|
Composition:
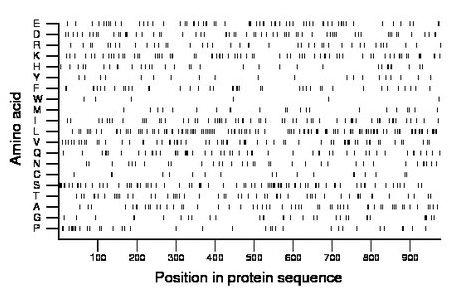
Amino acid Percentage Count Longest homopolymer A alanine 5.8 57 2 C cysteine 1.8 18 1 D aspartate 6.3 62 2 E glutamate 6.7 65 3 F phenylalanine 4.2 41 2 G glycine 3.4 33 2 H histidine 3.4 33 1 I isoleucine 6.1 60 2 K lysine 7.2 70 2 L leucine 12.0 117 3 M methionine 2.5 24 2 N asparagine 3.7 36 2 P proline 3.8 37 2 Q glutamine 5.3 52 2 R arginine 4.2 41 1 S serine 8.8 86 4 T threonine 5.6 55 2 V valine 6.6 64 3 W tryptophan 0.6 6 1 Y tyrosine 2.0 20 1 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 vacuolar protein sorting 54 isoform 1 VPS54 0.983 vacuolar protein sorting 54 isoform 2 CCDC132 0.010 coiled-coil domain containing 132 isoform a PMFBP1 0.008 polyamine modulated factor 1 binding protein 1 isof... PTPN3 0.007 protein tyrosine phosphatase, non-receptor type 3 i... PTPN3 0.007 protein tyrosine phosphatase, non-receptor type 3 i... PTPN3 0.007 protein tyrosine phosphatase, non-receptor type 3 i... PTPN3 0.006 protein tyrosine phosphatase, non-receptor type 3 i... PTPN3 0.006 protein tyrosine phosphatase, non-receptor type 3 i... PTPN3 0.006 protein tyrosine phosphatase, non-receptor type 3 i... LOC100130716 0.005 PREDICTED: similar to mucin 11 LOC100130716 0.005 PREDICTED: similar to Mucin-12 MUC12 0.005 PREDICTED: mucin 12 MUC12 0.005 PREDICTED: mucin 12 LOC100133714 0.005 PREDICTED: similar to Mucin-12, partial NEDD1 0.005 neural precursor cell expressed, developmentally do... NEDD1 0.005 neural precursor cell expressed, developmentally do... NEDD1 0.005 neural precursor cell expressed, developmentally dow... NEDD1 0.005 neural precursor cell expressed, developmentally do... MUC17 0.005 mucin 17 FAM21A 0.005 hypothetical protein LOC387680 KIF27 0.004 kinesin family member 27 HIP1R 0.004 huntingtin interacting protein-1-related SERPING1 0.004 serpin peptidase inhibitor, clade G, member 1 precur... SERPING1 0.004 serpin peptidase inhibitor, clade G, member 1 precur... LOC100294412 0.004 PREDICTED: similar to KIAA0655 protein FAM21B 0.004 hypothetical protein LOC55747 PMFBP1 0.004 polyamine modulated factor 1 binding protein 1 isof... GRIPAP1 0.004 GRIP1 associated protein 1 isoform 1 MUC12 0.004 PREDICTED: mucin 12, cell surface associatedHuman BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
