
| Name: THPO | Sequence: fasta or formatted (353aa) | NCBI GI: 4507493 | |
|
Description: thrombopoietin precursor
|
Referenced in: Platelets and Megakaryocytes
| ||
|
Composition:
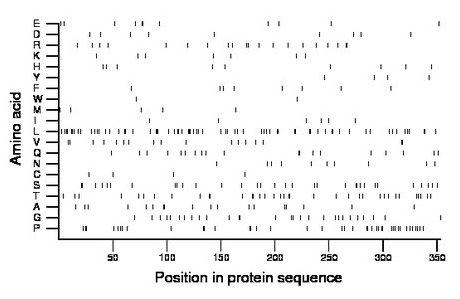
Amino acid Percentage Count Longest homopolymer A alanine 5.1 18 2 C cysteine 1.1 4 1 D aspartate 2.5 9 1 E glutamate 3.1 11 2 F phenylalanine 2.3 8 1 G glycine 7.9 28 2 H histidine 2.5 9 1 I isoleucine 1.7 6 1 K lysine 2.0 7 1 L leucine 19.3 68 3 M methionine 1.4 5 1 N asparagine 2.8 10 1 P proline 11.3 40 2 Q glutamine 5.4 19 2 R arginine 5.7 20 2 S serine 9.1 32 2 T threonine 10.2 36 2 V valine 4.8 17 2 W tryptophan 0.6 2 1 Y tyrosine 1.1 4 1 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 thrombopoietin precursor RICH2 0.032 Rho GTPase-activating protein RICH2 RBM12 0.029 RNA binding motif protein 12 RBM12 0.029 RNA binding motif protein 12 ACR 0.028 acrosin precursor LOC645529 0.026 PREDICTED: similar to Putative acrosin-like proteas... UBAP2L 0.025 ubiquitin associated protein 2-like isoform a [Homo... UBAP2L 0.025 ubiquitin associated protein 2-like isoform b [Homo... HOXA3 0.023 homeobox A3 isoform a HOXA3 0.023 homeobox A3 isoform a HTT 0.022 huntingtin FIP1L1 0.022 FIP1 like 1 isoform 1 WASF2 0.022 WAS protein family, member 2 SETD1B 0.022 SET domain containing 1B FLJ22184 0.021 PREDICTED: hypothetical protein FLJ22184 FLJ22184 0.021 PREDICTED: hypothetical protein LOC80164 FIP1L1 0.021 FIP1 like 1 isoform 2 PIAS3 0.021 protein inhibitor of activated STAT, 3 CPSF6 0.021 cleavage and polyadenylation specific factor 6, 68 ... NCCRP1 0.021 non-specific cytotoxic cell receptor protein 1 homol... MAPK1IP1L 0.021 MAPK-interacting and spindle-stabilizing protein [Ho... MUC2 0.021 mucin 2 precursor LOC100128719 0.021 PREDICTED: hypothetical protein, partial LOC100128719 0.021 PREDICTED: hypothetical protein, partial WASL 0.019 Wiskott-Aldrich syndrome gene-like protein PRRT2 0.019 proline-rich transmembrane protein 2 PHLDB1 0.019 pleckstrin homology-like domain, family B, member 1... PHLDB1 0.019 pleckstrin homology-like domain, family B, member 1... PHLDB1 0.019 pleckstrin homology-like domain, family B, member 1 ... LOC284297 0.019 hypothetical protein LOC284297Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
