
| Name: LOC649238 | Sequence: fasta or formatted (409aa) | NCBI GI: 169216732 | |
|
Description: PREDICTED: similar to hCG1645335
|
Referenced in: Additional Brain Proteins
| ||
Other entries for this name:
alt prot [409aa] PREDICTED: similar to hCG1645335 | |||
|
Composition:
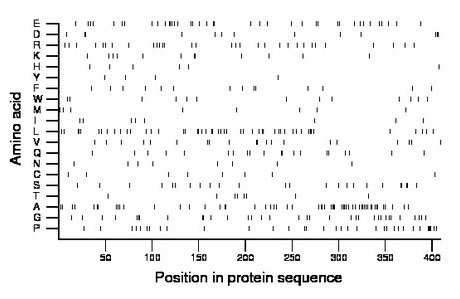
Amino acid Percentage Count Longest homopolymer A alanine 13.7 56 3 C cysteine 2.0 8 1 D aspartate 3.2 13 2 E glutamate 9.5 39 3 F phenylalanine 3.4 14 1 G glycine 8.3 34 2 H histidine 1.5 6 1 I isoleucine 2.0 8 1 K lysine 2.9 12 2 L leucine 12.0 49 2 M methionine 2.0 8 1 N asparagine 2.0 8 1 P proline 8.3 34 4 Q glutamine 4.9 20 2 R arginine 7.6 31 2 S serine 5.9 24 2 T threonine 2.0 8 1 V valine 4.9 20 2 W tryptophan 3.2 13 1 Y tyrosine 1.0 4 1 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary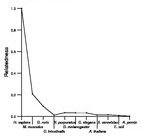 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 PREDICTED: similar to hCG1645335 LOC649238 0.995 PREDICTED: similar to hCG1645335 LOC649201 0.745 PREDICTED: similar to paraneoplastic antigen MA1 [H... LOC649201 0.744 PREDICTED: similar to paraneoplastic antigen MA1 [H... LOC649201 0.744 PREDICTED: similar to paraneoplastic antigen MA1 [H... LOC100287500 0.305 PREDICTED: hypothetical protein XP_002343900 PNMA6A 0.305 paraneoplastic antigen like 6A LOC100287466 0.302 PREDICTED: hypothetical protein XP_002343899 LOC100287428 0.302 PREDICTED: hypothetical protein XP_002343898 PNMA3 0.208 paraneoplastic cancer-testis-brain antigen PNMA5 0.198 paraneoplastic antigen like 5 PNMA5 0.198 paraneoplastic antigen like 5 PNMA5 0.198 paraneoplastic antigen like 5 LOC100128960 0.187 PREDICTED: similar to paraneoplastic antigen MA2 [H... LOC100128960 0.186 PREDICTED: similar to Putative paraneoplastic antig... PNMA1 0.174 paraneoplastic antigen MA1 MOAP1 0.155 modulator of apoptosis 1 PNMA2 0.139 paraneoplastic antigen MA2 ZCCHC12 0.106 zinc finger, CCHC domain containing 12 ZCCHC18 0.103 zinc finger, CCHC domain containing 18 PNMAL1 0.095 PNMA-like 1 isoform b PNMAL1 0.095 PNMA-like 1 isoform a PNMAL2 0.069 PNMA-like 2 TMPRSS13 0.021 transmembrane protease, serine 13 FAM48B1 0.020 hypothetical protein LOC100130302 MARCKS 0.017 myristoylated alanine-rich protein kinase C substra... FCAMR 0.017 Fc receptor, IgA, IgM, high affinity isoform 2 [Hom... KRTAP4-6 0.017 PREDICTED: keratin associated protein 4.6 UBQLN4 0.017 ataxin-1 ubiquitin-like interacting protein C8orf47 0.017 hypothetical protein LOC203111Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
