
| Name: SHC4 | Sequence: fasta or formatted (630aa) | NCBI GI: 222446609 | |
|
Description: rai-like protein
|
Referenced in: Non-Receptor Tyrosine Kinase Pathways
| ||
|
Composition:
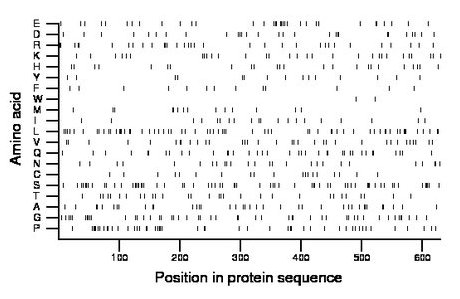
Amino acid Percentage Count Longest homopolymer A alanine 6.0 38 2 C cysteine 3.0 19 1 D aspartate 4.4 28 2 E glutamate 5.2 33 2 F phenylalanine 2.1 13 1 G glycine 7.8 49 2 H histidine 4.0 25 2 I isoleucine 3.5 22 2 K lysine 4.9 31 2 L leucine 10.2 64 3 M methionine 2.5 16 1 N asparagine 4.1 26 2 P proline 7.8 49 3 Q glutamine 5.9 37 2 R arginine 5.2 33 1 S serine 10.2 64 2 T threonine 4.9 31 2 V valine 5.4 34 1 W tryptophan 0.3 2 1 Y tyrosine 2.5 16 2 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 rai-like protein SHC1 0.354 SHC (Src homology 2 domain containing) transforming... SHC1 0.354 SHC (Src homology 2 domain containing) transforming... SHC3 0.338 src homology 2 domain-containing transforming prote... SHC1 0.320 SHC (Src homology 2 domain containing) transforming ... SHC1 0.319 SHC (Src homology 2 domain containing) transforming... SHC2 0.276 SHC (Src homology 2 domain containing) transforming... LOC100291393 0.081 PREDICTED: hypothetical protein XP_002347924 FER 0.037 fer (fps/fes related) tyrosine kinase BCAR3 0.027 breast cancer antiestrogen resistance 3 SH2B2 0.026 SH2B adaptor protein 2 SH2D3C 0.025 SH2 domain containing 3C isoform e SH2D3C 0.025 SH2 domain containing 3C isoform d SH2D3C 0.025 SH2 domain containing 3C isoform b SH2D3C 0.025 SH2 domain containing 3C isoform a FES 0.023 feline sarcoma oncogene isoform 2 FES 0.023 feline sarcoma oncogene isoform 1 ABL2 0.021 arg tyrosine kinase isoform c ABL2 0.021 arg tyrosine kinase isoform b ABL2 0.021 arg tyrosine kinase isoform e ABL2 0.021 arg tyrosine kinase isoform d ABL2 0.021 arg tyrosine kinase isoform a SH2B1 0.021 SH2B adaptor protein 1 isoform 1 SH2B1 0.021 SH2B adaptor protein 1 isoform 2 SH2B1 0.021 SH2B adaptor protein 1 isoform 3 SH2B1 0.021 SH2B adaptor protein 1 isoform 2 SH2B1 0.021 SH2B adaptor protein 1 isoform 2 APBA1 0.021 amyloid beta A4 precursor protein-binding, family A,... SHB 0.021 Src homology 2 domain containing adaptor protein B ... SHF 0.021 Src homology 2 domain containing FHuman BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
