
| Name: LOC728637 | Sequence: fasta or formatted (213aa) | NCBI GI: 239748651 | |
|
Description: PREDICTED: similar to rCG33261
| Not currently referenced in the text | ||
Other entries for this name:
alt mRNA {213aa} PREDICTED: similar to rCG33261 alt mRNA {213aa} PREDICTED: similar to rCG33261 | |||
|
Composition:
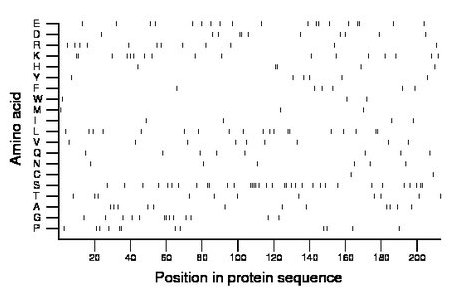
Amino acid Percentage Count Longest homopolymer A alanine 5.6 12 1 C cysteine 0.9 2 1 D aspartate 5.2 11 2 E glutamate 8.9 19 2 F phenylalanine 2.8 6 1 G glycine 6.1 13 2 H histidine 2.3 5 2 I isoleucine 1.9 4 1 K lysine 8.0 17 2 L leucine 8.9 19 2 M methionine 1.4 3 1 N asparagine 2.8 6 1 P proline 5.6 12 2 Q glutamine 3.8 8 1 R arginine 5.6 12 1 S serine 16.4 35 4 T threonine 5.2 11 1 V valine 4.2 9 1 W tryptophan 1.4 3 1 Y tyrosine 2.8 6 1 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 PREDICTED: similar to rCG33261 LOC728637 1.000 PREDICTED: similar to rCG33261 LOC728637 1.000 PREDICTED: similar to rCG33261 TNRC18 0.035 trinucleotide repeat containing 18 TTN 0.025 titin isoform N2-A UNC84B 0.022 unc-84 homolog B PDZD4 0.020 PDZ domain containing 4 NEFH 0.020 neurofilament, heavy polypeptide 200kDa MICAL1 0.020 microtubule associated monoxygenase, calponin and L... MICAL1 0.020 microtubule associated monoxygenase, calponin and L... HNRNPUL2 0.017 heterogeneous nuclear ribonucleoprotein U-like 2 [H... C22orf42 0.017 chromosome 22 open reading frame 42 TBX2 0.017 T-box 2 STK36 0.017 serine/threonine kinase 36 DMP1 0.015 dentin matrix acidic phosphoprotein 1 isoform 1 precu... DMP1 0.015 dentin matrix acidic phosphoprotein 1 isoform 2 pre... LOC100293861 0.015 PREDICTED: hypothetical protein LOC100291362 0.015 PREDICTED: hypothetical protein XP_002346476 LOC100286920 0.015 PREDICTED: hypothetical protein XP_002342339 TBC1D9B 0.015 TBC1 domain family, member 9B (with GRAM domain) iso... TBC1D9B 0.015 TBC1 domain family, member 9B (with GRAM domain) iso... PSD3 0.012 ADP-ribosylation factor guanine nucleotide factor 6... C17orf76 0.012 hypothetical protein LOC388341 isoform 1 RTN3 0.012 reticulon 3 isoform b JPH2 0.012 junctophilin 2 isoform 1 RBM15B 0.012 RNA binding motif protein 15B CDC42EP1 0.012 CDC42 effector protein 1 NCL 0.012 nucleolin SETD1A 0.012 SET domain containing 1A AATK 0.012 apoptosis-associated tyrosine kinaseHuman BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
