
| Name: LOC100292771 | Sequence: fasta or formatted (126aa) | NCBI GI: 239753604 | |
|
Description: PREDICTED: hypothetical protein
| Not currently referenced in the text | ||
|
Composition:
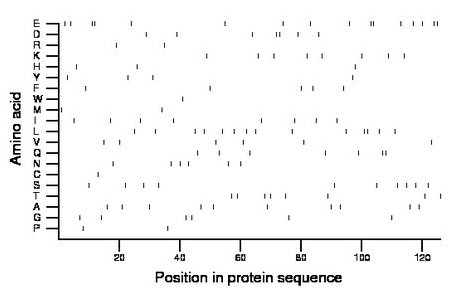
Amino acid Percentage Count Longest homopolymer A alanine 7.9 10 1 C cysteine 0.8 1 1 D aspartate 5.6 7 2 E glutamate 12.7 16 2 F phenylalanine 4.0 5 1 G glycine 4.8 6 1 H histidine 2.4 3 1 I isoleucine 6.3 8 1 K lysine 6.3 8 1 L leucine 11.1 14 2 M methionine 1.6 2 1 N asparagine 4.8 6 1 P proline 1.6 2 1 Q glutamine 5.6 7 2 R arginine 1.6 2 1 S serine 7.9 10 1 T threonine 6.3 8 1 V valine 4.8 6 1 W tryptophan 0.8 1 1 Y tyrosine 3.2 4 1 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary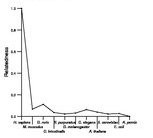 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 PREDICTED: hypothetical protein LOC100287790 1.000 PREDICTED: hypothetical protein LOC100287790 1.000 PREDICTED: hypothetical protein XP_002342430 FTH1 0.071 ferritin, heavy polypeptide 1 FTMT 0.058 ferritin mitochondrial FTL 0.040 ferritin, light polypeptide LOC139431 0.040 PREDICTED: similar to hCG1799751 LOC139431 0.040 PREDICTED: similar to hCG1799751 LOC139431 0.040 PREDICTED: hypothetical protein LOC139431 BSDC1 0.022 BSD domain containing 1 isoform d BSDC1 0.022 BSD domain containing 1 isoform c BSDC1 0.022 BSD domain containing 1 isoform a BSDC1 0.022 BSD domain containing 1 isoform b FTHL17 0.022 ferritin, heavy polypeptide-like 17 MTUS1 0.018 mitochondrial tumor suppressor 1 isoform 5 MTUS1 0.018 mitochondrial tumor suppressor 1 isoform 1 MTUS1 0.018 mitochondrial tumor suppressor 1 isoform 2 MTUS1 0.018 mitochondrial tumor suppressor 1 isoform 4 PAPOLB 0.018 poly(A) polymerase beta (testis specific) PAPOLA 0.018 poly(A) polymerase alpha USP25 0.013 ubiquitin specific peptidase 25 SCN4B 0.013 sodium channel, voltage-gated, type IV, beta isoform... PLEC1 0.013 plectin 1 isoform 11 PLEC1 0.013 plectin 1 isoform 10 PLEC1 0.013 plectin 1 isoform 8 PLEC1 0.013 plectin 1 isoform 7 PLEC1 0.013 plectin 1 isoform 6 PLEC1 0.013 plectin 1 isoform 3 PLEC1 0.013 plectin 1 isoform 2 PLEC1 0.013 plectin 1 isoform 1Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
