
| Name: PPAT | Sequence: fasta or formatted (517aa) | NCBI GI: 29570798 | |
|
Description: phosphoribosyl pyrophosphate amidotransferase proprotein
|
Referenced in: Purine Biosynthesis and Catabolism
| ||
|
Composition:
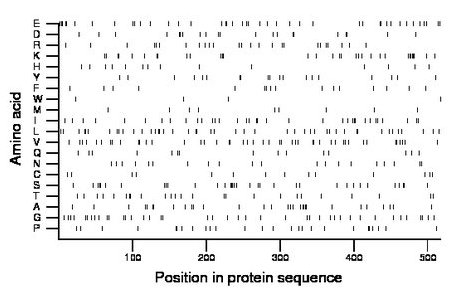
Amino acid Percentage Count Longest homopolymer A alanine 5.8 30 2 C cysteine 2.7 14 1 D aspartate 4.1 21 2 E glutamate 8.1 42 2 F phenylalanine 3.1 16 1 G glycine 7.9 41 1 H histidine 2.7 14 1 I isoleucine 6.8 35 2 K lysine 6.4 33 2 L leucine 8.9 46 2 M methionine 1.9 10 1 N asparagine 3.9 20 1 P proline 5.0 26 2 Q glutamine 3.1 16 2 R arginine 4.8 25 1 S serine 6.8 35 2 T threonine 4.8 25 2 V valine 8.7 45 2 W tryptophan 0.8 4 1 Y tyrosine 3.7 19 2 |
Comparative genomics:
Search single species RefSeq proteins at NCBI
Search summary 
Figure data | ||
Related human proteins:Protein Relative score Description Self-match 1.000 phosphoribosyl pyrophosphate amidotransferase propro... GFPT2 0.036 glutamine-fructose-6-phosphate transaminase 2 GFPT1 0.031 glucosamine-fructose-6-phosphate aminotransferase [... MKI67 0.007 antigen identified by monoclonal antibody Ki-67 iso... MKI67 0.007 antigen identified by monoclonal antibody Ki-67 iso... HPRT1 0.006 hypoxanthine phosphoribosyltransferase 1 NOS2 0.006 nitric oxide synthase 2A FOXD2 0.006 forkhead box D2 MUC16 0.005 mucin 16 GUSB 0.005 glucuronidase, beta RNF2 0.004 ring finger protein 2 HMGXB3 0.004 HMG box domain containing 3Human BLASTP results (used to prepare the table) | |||
Gene descriptions are from NCBI RefSeq. Search results were obtained with NCBI BLAST and RefSeq entries. When identical proteins are present, the self-match may not be listed first in BLASTP output. In such cases, the table above has been reordered to place it first.
See About the Figures for the scoring system used in the figure above right. The same scoring system was used in the table of BLASTP results.
Guide to the Human Genome
Copyright © 2010 by Stewart Scherer. All rights reserved.
